Computational methods for region-based analysis of epigenome signals
Nathan Sheffield, PhD

outline
The (epi)genome revolution
Epigenome tools
|
|
20%
40%
40%
|
Project organization
◁ Questions ▷
The genome revolution

A revolution driven by DNA sequencing technology
Epigenomics is the study of the chemical modification and physical conformation of cellular DNA and bound proteins


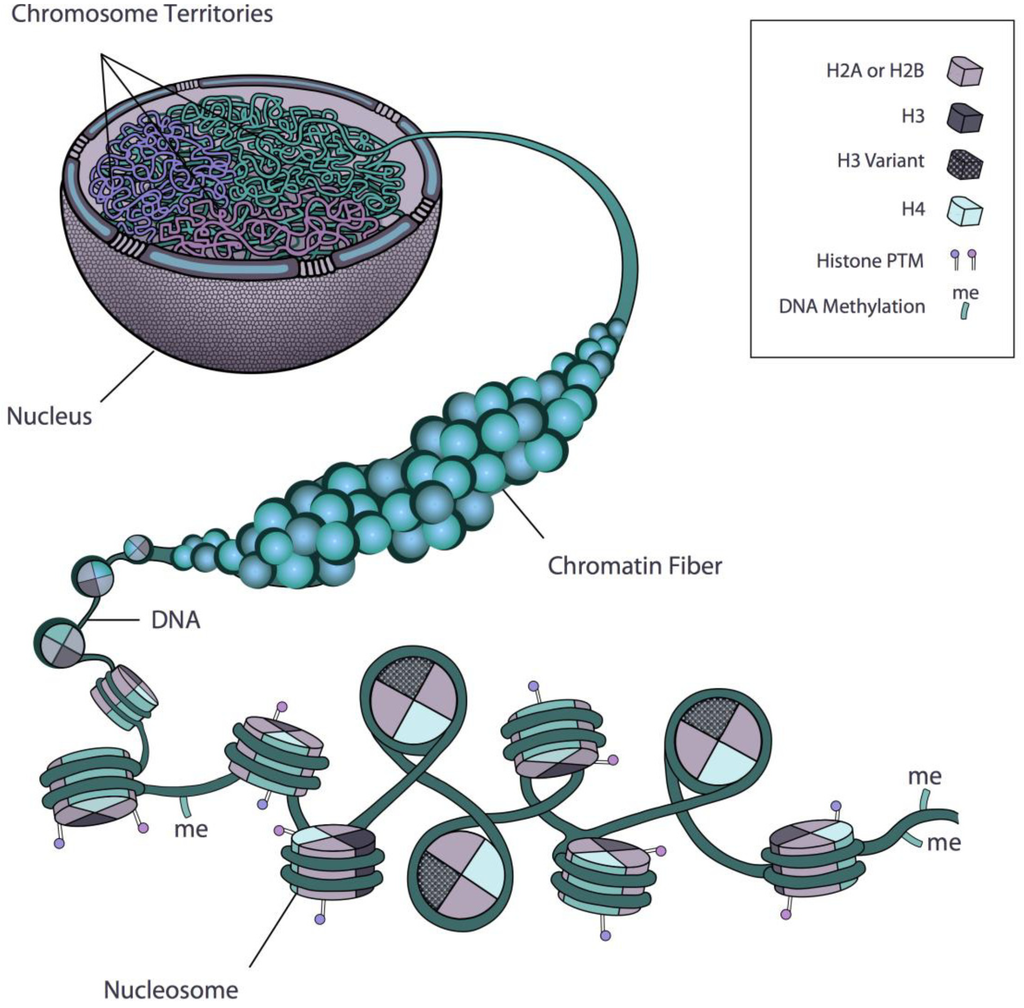
Rosa et al. 2013
- Histone modification:
ChIP-seq - DNA methylation:
Bisulfite-seq - Chromatin accessibility:
ATAC-seq
The Sequence Read Archive is growing
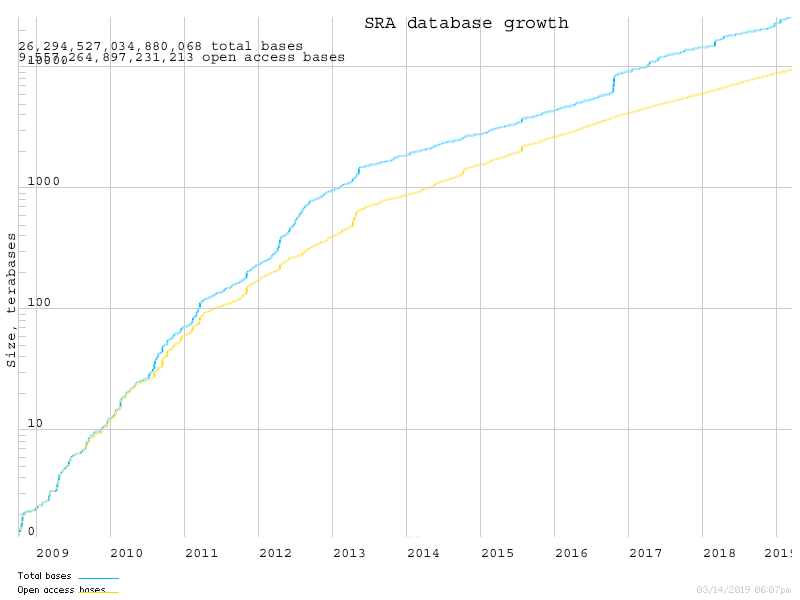 https://www.ncbi.nlm.nih.gov/sra/docs/sragrowth/
https://www.ncbi.nlm.nih.gov/sra/docs/sragrowth/
abundant
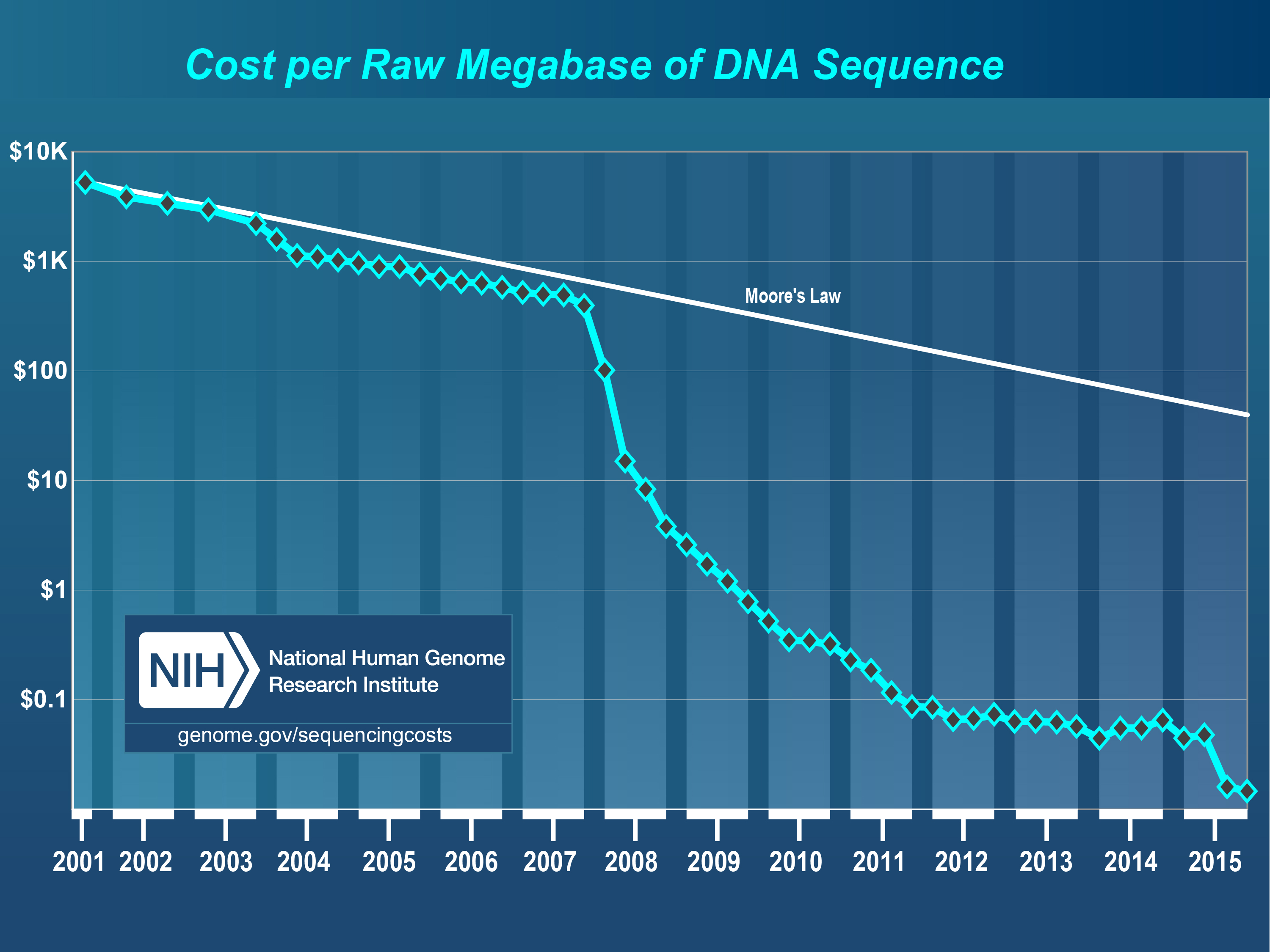

available


powerful




First step in bioinformatics analysis:

pipeline
pipeline

Papers with
"bioinformatics pipeline"
in title



Data munging


Then, downstream tools need a different organization

What if?


Microwave syndrome
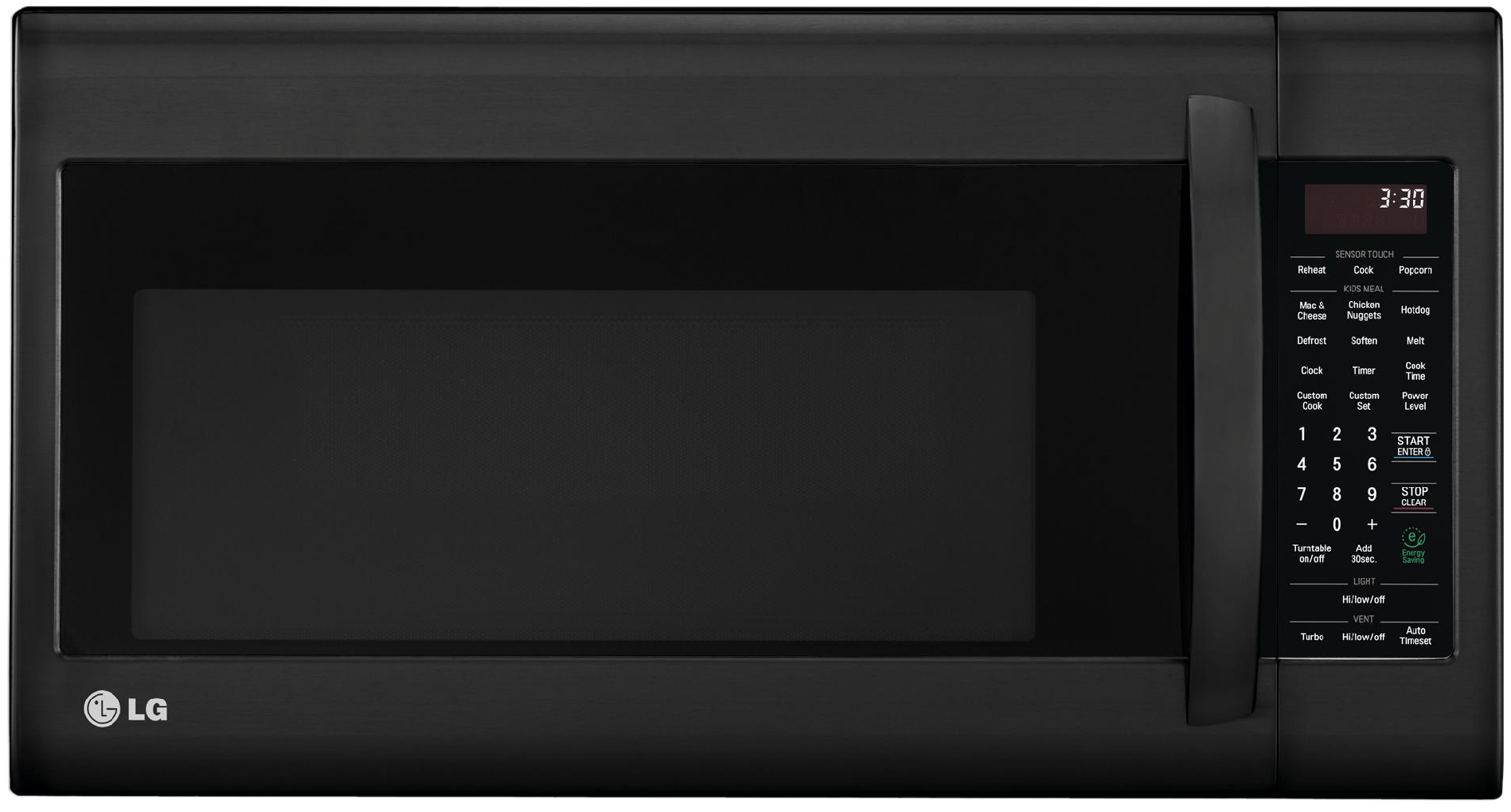
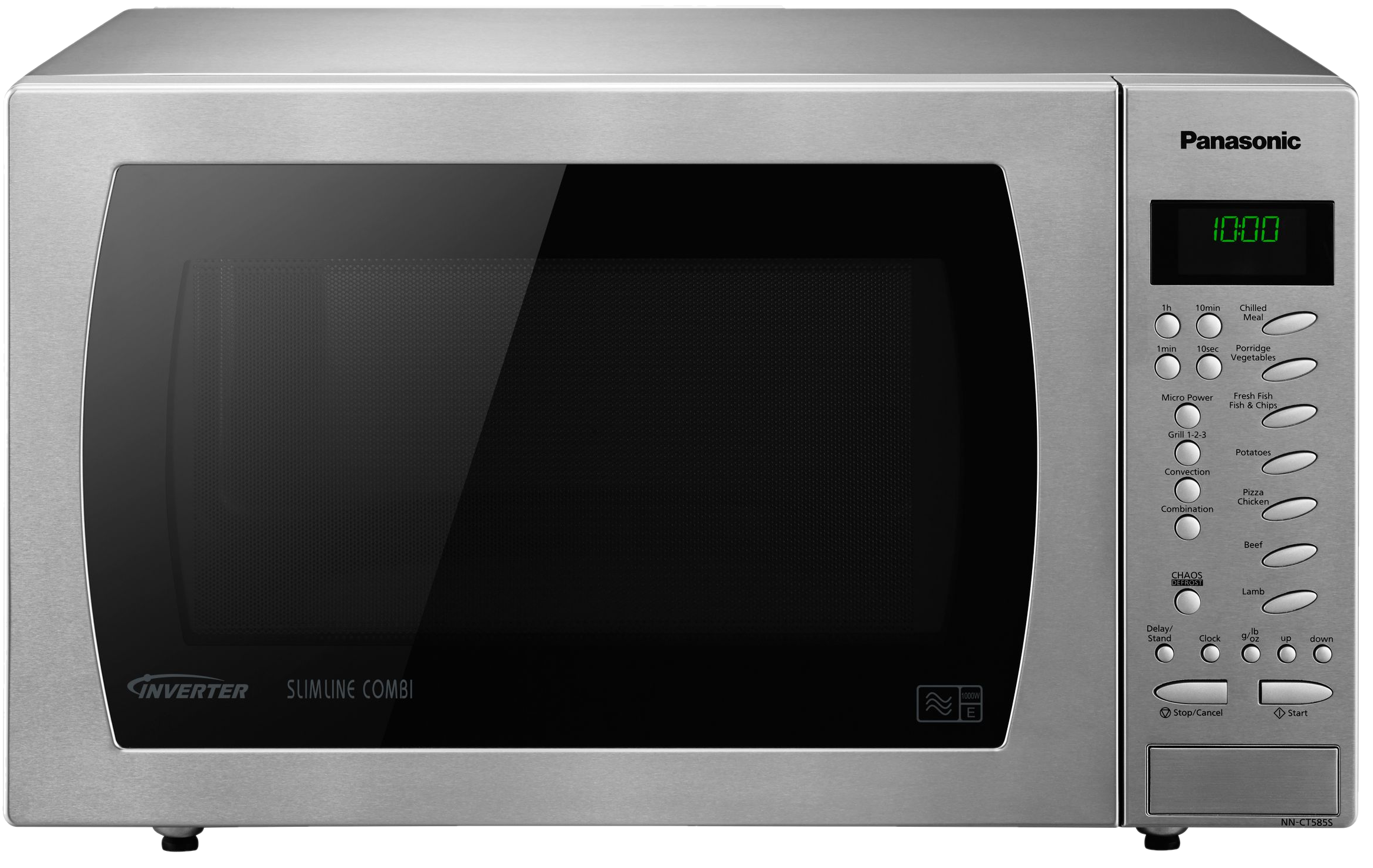



In user interface design, prioritizing easy access to integrated functions over their individual components.







The UNIX philosophy

[T]he power of a system comes more from the relationships among programs than from the programs themselves.
Many UNIX programs do quite trivial tasks in isolation, but, combined with other programs, become general and useful tools.
- Kernighan and Pike, The UNIX Programming Environment (1983, p. viii)
Many UNIX programs do quite trivial tasks in isolation, but, combined with other programs, become general and useful tools.
- Kernighan and Pike, The UNIX Programming Environment (1983, p. viii)







Problem

Solution

Problem

Solution

PEP: Portable Encapsulated Projects

 PEP format
PEP format
sample_name,protocol,organism,input_file
frog_0h,RNA-seq,frog,/path/to/frog0.gz
frog_1h,RNA-seq,frog,/path/to/frog1.gz
frog_2h,RNA-seq,frog,/path/to/frog2.gz
frog_3h,RNA-seq,frog,/path/to/frog3.gz
 PEP format
PEP format
sample_name,protocol,organism,input_file
frog_0h,RNA-seq,frog,/path/to/frog0.gz
frog_1h,RNA-seq,frog,/path/to/frog1.gz
frog_2h,RNA-seq,frog,/path/to/frog2.gz
frog_3h,RNA-seq,frog,/path/to/frog3.gz
sample_table: /path/to/samples.csv
output_dir: /path/to/output/folder
other_variable: value
Add programmatic sample and project modifiers.
Automatically build new sample attributes from existing attributes.
Without derived attribute:
| sample_name | t | protocol | organism | input_file |
| ------------- | ---- | :-------------: | -------- | ---------------------- |
| frog_0h | 0 | RNA-seq | frog | /path/to/frog0.gz |
| frog_1h | 1 | RNA-seq | frog | /path/to/frog1.gz |
| frog_2h | 2 | RNA-seq | frog | /path/to/frog2.gz |
| frog_3h | 3 | RNA-seq | frog | /path/to/frog3.gz |
Using derived attribute:
| sample_name | t | protocol | organism | input_file |
| ------------- | ---- | :-------------: | -------- | ---------------------- |
| frog_0h | 0 | RNA-seq | frog | my_samples |
| frog_1h | 1 | RNA-seq | frog | my_samples |
| frog_2h | 2 | RNA-seq | frog | my_samples |
| frog_3h | 3 | RNA-seq | frog | my_samples |
| crab_0h | 0 | RNA-seq | crab | your_samples |
| crab_3h | 3 | RNA-seq | crab | your_samples |
| sample_name | t | protocol | organism | input_file |
| ------------- | ---- | :-------------: | -------- | ---------------------- |
| frog_0h | 0 | RNA-seq | frog | my_samples |
| frog_1h | 1 | RNA-seq | frog | my_samples |
| frog_2h | 2 | RNA-seq | frog | my_samples |
| frog_3h | 3 | RNA-seq | frog | my_samples |
| crab_0h | 0 | RNA-seq | crab | your_samples |
| crab_3h | 3 | RNA-seq | crab | your_samples |
Project config file:
sample_modifiers:
derive:
attributes: [input_file]
sources:
my_samples: "/path/to/my/samples/{organism}_{t}h.gz"
your_samples: "/path/to/your/samples/{organism}_{t}h.gz"Benefit: Enables distributed files, portability
Add new sample attributes conditioned on values of existing attributes
Before:
| sample_name | protocol | organism |
| ------------- | :-------------: | -------- |
| human_1 | RNA-seq | human |
| human_2 | RNA-seq | human |
| human_3 | RNA-seq | human |
| mouse_1 | RNA-seq | mouse |
After:
| sample_name | protocol | organism | genome |
| ------------- | :-------------: | -------- | ------ |
| human_1 | RNA-seq | human | hg38 |
| human_2 | RNA-seq | human | hg38 |
| human_3 | RNA-seq | human | hg38 |
| mouse_1 | RNA-seq | mouse | mm10 |
| sample_name | protocol | organism |
| ------------- | :-------------: | -------- |
| human_1 | RNA-seq | human |
| human_2 | RNA-seq | human |
| human_3 | RNA-seq | human |
| mouse_1 | RNA-seq | mouse |
Project config file:
sample_modifiers:
imply:
- if:
organism: human
then:
genome: hg38
- if:
organism: mouse
then:
genome: mm10Benefit: Divides project from sample metadata
Define activatable project attributes.
project_modifiers:
amendments:
diverse:
metadata:
sample_annotation: psa_rrbs_diverse.csv
cancer:
metadata:
sample_annotation: psa_rrbs_intracancer.csvBenefit: Defines multiple similar projects in a single file


Locus Overlap Analysis












If subject list has no containment,
identifying overlaps is fast

binary search on start intervals, followed by backward steps:


The problem arises with contained interval overlaps


How can we improve efficiency
without guaranteeing no containment?
Many approaches to solve the 'containment' issue:
- Nested Containment Lists (GRanges) [@Alekseyenko2007; @Aboyoun2012] - R-trees (bedtools) [@Kent2002; @Quinlan2010], Augmented interval trees [@Cormen2001] These methods try to structure the data to provide non-containment guaranteesMethods provide non-containment guarantees
R-trees
Annotates tree nodes with a minimum bounding rectangle of elements. A query that does not intersect the bounding rectangle will not intersect any child element.Nested Containment Lists

Augmented Interval List
1. Augment the list with the running maximum *end* value. *solves the problem for lowly-contained lists* 2. Decompose the list to minimize containment. *extends the solution to highly-contained lists*Augment with the running maximum end value, `maxE`
Provides a local guarantee of no containment.
AIList works on contained lists


But long containment runs are problematic


Decompose long runs with constant `maxE`

Performance
- How does the `maxE` minimum run length affect performance?
- How does it compare to existing approaches?
- How does it scale with increasing size of subject?
Datasets

How does the `maxE` minimum run length affect performance?

How does it compare to existing approaches?

How does it scale with increasing size of subject?

Conclusion
- Augmented Interval Lists add the maximum running end value to a list of intervals
- The data structure is simpler than other methods
- AILists improve performance, particularly in highly contained interval sets
Conclusion
Thank You
Sheffield lab
John Lawson
Vince Reuter
Jason Smith
Jianglin Feng
Michal Stolarczyk
Aaron Gu
Ognen Duzlevski
SOM research computing
Pete Nagraj
Neal Magee
Funding:





