Organizing large-scale biological data around standardized projects
Nathan Sheffield, PhD

abundant
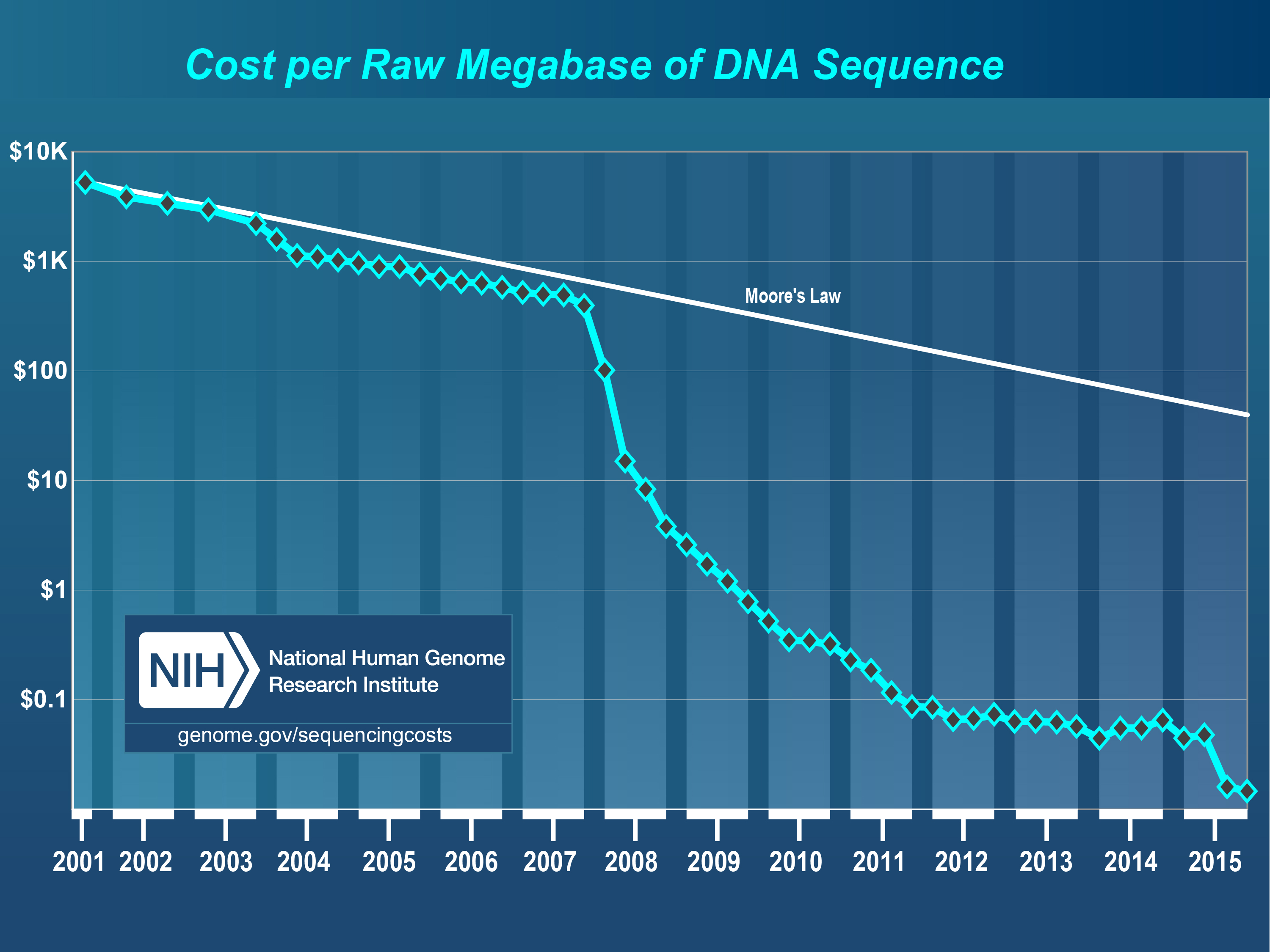

available


powerful




First step in bioinformatics analysis:

pipeline
pipeline

Papers with
"bioinformatics pipeline"
in title



Data munging


Then, downstream tools need a different organization

What if?


Why is this hard to do?
Because of microwave syndrome....
Because of microwave syndrome....
Microwave syndrome



In user interface design, prioritizing easy access to integrated functions over their individual components.







The UNIX philosophy

[T]he power of a system comes more from the relationships among programs than from the programs themselves.
Many UNIX programs do quite trivial tasks in isolation, but, combined with other programs, become general and useful tools.
- Kernighan and Pike, The UNIX Programming Environment (1983, p. viii)
Many UNIX programs do quite trivial tasks in isolation, but, combined with other programs, become general and useful tools.
- Kernighan and Pike, The UNIX Programming Environment (1983, p. viii)







Problem

Solution


PEP: Portable Encapsulated Projects

 PEP format
PEP format
sample_name,protocol,organism,input_file
frog_0h,RNA-seq,frog,/path/to/frog0.gz
frog_1h,RNA-seq,frog,/path/to/frog1.gz
frog_2h,RNA-seq,frog,/path/to/frog2.gz
frog_3h,RNA-seq,frog,/path/to/frog3.gz
 PEP format
PEP format
sample_name,protocol,organism,input_file
frog_0h,RNA-seq,frog,/path/to/frog0.gz
frog_1h,RNA-seq,frog,/path/to/frog1.gz
frog_2h,RNA-seq,frog,/path/to/frog2.gz
frog_3h,RNA-seq,frog,/path/to/frog3.gz
sample_table: /path/to/samples.csv
output_dir: /path/to/output/folder
other_variable: value
Add programmatic sample and project modifiers.
Automatically build new sample attributes from existing attributes.
Without derived attribute:
| sample_name | t | protocol | organism | input_file |
| ------------- | ---- | :-------------: | -------- | ---------------------- |
| frog_0h | 0 | RNA-seq | frog | /path/to/frog0.gz |
| frog_1h | 1 | RNA-seq | frog | /path/to/frog1.gz |
| frog_2h | 2 | RNA-seq | frog | /path/to/frog2.gz |
| frog_3h | 3 | RNA-seq | frog | /path/to/frog3.gz |
Using derived attribute:
| sample_name | t | protocol | organism | input_file |
| ------------- | ---- | :-------------: | -------- | ---------------------- |
| frog_0h | 0 | RNA-seq | frog | my_samples |
| frog_1h | 1 | RNA-seq | frog | my_samples |
| frog_2h | 2 | RNA-seq | frog | my_samples |
| frog_3h | 3 | RNA-seq | frog | my_samples |
| crab_0h | 0 | RNA-seq | crab | your_samples |
| crab_3h | 3 | RNA-seq | crab | your_samples |
| sample_name | t | protocol | organism | input_file |
| ------------- | ---- | :-------------: | -------- | ---------------------- |
| frog_0h | 0 | RNA-seq | frog | my_samples |
| frog_1h | 1 | RNA-seq | frog | my_samples |
| frog_2h | 2 | RNA-seq | frog | my_samples |
| frog_3h | 3 | RNA-seq | frog | my_samples |
| crab_0h | 0 | RNA-seq | crab | your_samples |
| crab_3h | 3 | RNA-seq | crab | your_samples |
Project config file:
sample_modifiers:
derive:
attributes: [input_file]
sources:
my_samples: "/path/to/my/samples/{organism}_{t}h.gz"
your_samples: "/path/to/your/samples/{organism}_{t}h.gz"Benefit: Enables distributed files, portability
Add new sample attributes conditioned on values of existing attributes
Before:
| sample_name | protocol | organism |
| ------------- | :-------------: | -------- |
| human_1 | RNA-seq | human |
| human_2 | RNA-seq | human |
| human_3 | RNA-seq | human |
| mouse_1 | RNA-seq | mouse |
After:
| sample_name | protocol | organism | genome |
| ------------- | :-------------: | -------- | ------ |
| human_1 | RNA-seq | human | hg38 |
| human_2 | RNA-seq | human | hg38 |
| human_3 | RNA-seq | human | hg38 |
| mouse_1 | RNA-seq | mouse | mm10 |
| sample_name | protocol | organism |
| ------------- | :-------------: | -------- |
| human_1 | RNA-seq | human |
| human_2 | RNA-seq | human |
| human_3 | RNA-seq | human |
| mouse_1 | RNA-seq | mouse |
Project config file:
sample_modifiers:
imply:
- if:
organism: human
then:
genome: hg38
- if:
organism: mouse
then:
genome: mm10Benefit: Divides project from sample metadata
Define activatable project attributes.
project_modifiers:
amendments:
diverse:
metadata:
sample_annotation: psa_rrbs_diverse.csv
cancer:
metadata:
sample_annotation: psa_rrbs_intracancer.csvBenefit: Defines multiple similar projects in a single file
peppy package 

import peppy
prj = Project("pep_config.yaml")
samples = prj.get_samples()
for sample in samples:
print(sample.name)
# do further analysis to each sample
pepr package 

library("pepr")
prj = pepr::Project("pep_config.yaml")
samples = pepr::pepSamples(prj)
for (sample in samples) {
message(pepr::sampleName(sample))
# do further analysis to each sample
}

Looper
Deploys pipelines across samples by connectingsamples to any command-line tool

protocol_mappings:
RNA-seq: rna-seq
pipelines:
rna-seq:
name: RNA-seq_pipeline
path: path/to/rna-seq.py
arguments:
"--option1": sample_attribute
"--option2": sample_attribute2Looper features
Run your entire project with one line:
looper run project_config.yamlprotocol_mappings:
RRBS: rrbs
WGBS: wgbs
EG: wgbs.py
SMART-seq: rnaBitSeq -f; rnaTopHat -f
ATAC-SEQ: atacseq
DNase-seq: atacseq
CHIP-SEQ: chipseqpipeline_key:
name: pipeline_name
arguments:
"--option" : value
resources:
default:
file_size: "0"
cores: "2"
mem: "6000"
time: "01:00:00"
large_input:
file_size: "2000"
cores: "4"
mem: "12000"
time: "08:00:00"compute:
slurm:
submission_template: templates/slurm_template.sub
submission_command: sbatch
localhost:
submission_template: templates/localhost_template.sub
submission_command: shAdjust compute package on-the-fly:
> looper run project_config.yaml --compute localhostLooper only submits jobs for samples not already flagged as running, completed, or failed.
looper check project_config.yamllooper summarize project_config.yamlConclusion
- PEP format is a novel approach to standardize projects.
- Initial tools like
geofetchandlooperbuild PEP projects and connect them to pipelines - Python and R packages provide a universal interface to PEP metadata for tools and analysis
Thank You
