ATAC-seq pipeline processing
Nathan Sheffield, PhD


PEPATAC strengths
Modular system
Prealignments
Prealignments
Flexibility and portability
Outputs
Outputs
$ /pipelines/pepatac.py -h
usage: pepatac.py [-h] [-R] [-N] [-D] [-F] [-C CONFIG_FILE]
[-O PARENT_OUTPUT_FOLDER] [-M MEMORY_LIMIT]
[-P NUMBER_OF_CORES] -S SAMPLE_NAME -I INPUT_FILES
[INPUT_FILES ...] [-I2 [INPUT_FILES2 [INPUT_FILES2 ...]]] -G
GENOME_ASSEMBLY [-Q SINGLE_OR_PAIRED] [-gs GENOME_SIZE]
[--frip-ref-peaks FRIP_REF_PEAKS] [--TSS-name TSS_NAME]
[--anno-name ANNO_NAME] [--keep]
[--peak-caller {fseq,macs2}]
[--trimmer {trimmomatic,skewer}]
[--prealignments PREALIGNMENTS [PREALIGNMENTS ...]] [-V]
PEPATAC version 0.7.3
optional arguments:
-h, --help show this help message and exit
-R, --recover Overwrite locks to recover from previous failed run
-N, --new-start Overwrite all results to start a fresh run
-D, --dirty Don't auto-delete intermediate files
-F, --force-follow Always run 'follow' commands
-C CONFIG_FILE, --config CONFIG_FILE
Pipeline configuration file (YAML). Relative paths are
with respect to the pipeline script.
-O PARENT_OUTPUT_FOLDER, --output-parent PARENT_OUTPUT_FOLDER
Parent output directory of project
-M MEMORY_LIMIT, --mem MEMORY_LIMIT
Memory limit (in Mb) for processes accepting such
-P NUMBER_OF_CORES, --cores NUMBER_OF_CORES
Number of cores for parallelized processes
-I2 [INPUT_FILES2 [INPUT_FILES2 ...]], --input2 [INPUT_FILES2 [INPUT_FILES2 ...]]
Secondary input files, such as read2
-Q SINGLE_OR_PAIRED, --single-or-paired SINGLE_OR_PAIRED
Single- or paired-end sequencing protocol
-gs GENOME_SIZE, --genome-size GENOME_SIZE
genome size for MACS2
--frip-ref-peaks FRIP_REF_PEAKS
Reference peak set for calculating FRiP
--TSS-name TSS_NAME Name of TSS annotation file
--anno-name ANNO_NAME
Name of reference bed file for calculating FRiF
--keep Keep prealignment BAM files
--peak-caller {fseq,macs2}
Name of peak caller
--trimmer {trimmomatic,pyadapt,skewer}
Name of read trimming program
--prealignments PREALIGNMENTS [PREALIGNMENTS ...]
Space-delimited list of reference genomes to align to
before primary alignment.
-V, --version show program's version number and exit
required named arguments:
-S SAMPLE_NAME, --sample-name SAMPLE_NAME
Name for sample to run
-I INPUT_FILES [INPUT_FILES ...], --input INPUT_FILES [INPUT_FILES ...]
One or more primary input files
-G GENOME_ASSEMBLY, --genome GENOME_ASSEMBLY
Identifier for genome assembly


PEP specification for sample metadata
1. Configuration file:config.yaml
pep_version: 2.0.0
sample_table: "path/to/sample_table.csv"
sample_table.csv:
"sample_name", "protocol", "file"
"frog_1", "ATAC-seq", "frog1.fq.gz"
"frog_2", "ATAC-seq", "frog2.fq.gz"
"frog_3", "ATAC-seq", "frog3.fq.gz"
"frog_4", "ATAC-seq", "frog4.fq.gz"
MapReduce or Scatter/Gather
1. Map/Scatter PEPATAC across individual sampleslooper run config.yaml
looper runp config.yaml
PEPATAC strengths
Modular system
Prealignments
Prealignments
Flexibility and portability
Outputs
Outputs
Nuclear-mitochondrial DNA (NuMts) confuse aligners














PEPATAC strengths
Modular system
Prealignments
Prealignments
Flexibility and portability
Outputs
Outputs
Flexibility and Portability
pepatac.yaml# basic tools
tools: # absolute paths to required tools
java: java
python: python
samtools: samtools
bedtools: bedtools
bowtie2: bowtie2
fastqc: fastqc
macs2: macs2
picard: ${PICARD}
skewer: skewer
perl: perl
# ucsc tools
bedGraphToBigWig: bedGraphToBigWig
wigToBigWig: wigToBigWig
bigWigCat: bigWigCat
bedSort: bedSort
bedToBigBed: bedToBigBed
# optional tools
fseq: fseq
trimmo: ${TRIMMOMATIC}
Rscript: Rscript
# user configure
resources:
genomes: ${GENOMES}
adapters: null # Set to null to use default adapters
parameters: # parameters passed to bioinformatic tools
samtools:
q: 10
macs2:
f: BED
q: 0.01
shift: 0
fseq:
of: npf # narrowPeak as output format
l: 600 # feature length
t: 4.0 # "threshold" (standard deviations)
s: 1 # wiggle track stepPEPATAC strengths
Modular system
Prealignments
Prealignments
Flexibility and portability
Outputs
Outputs
Output

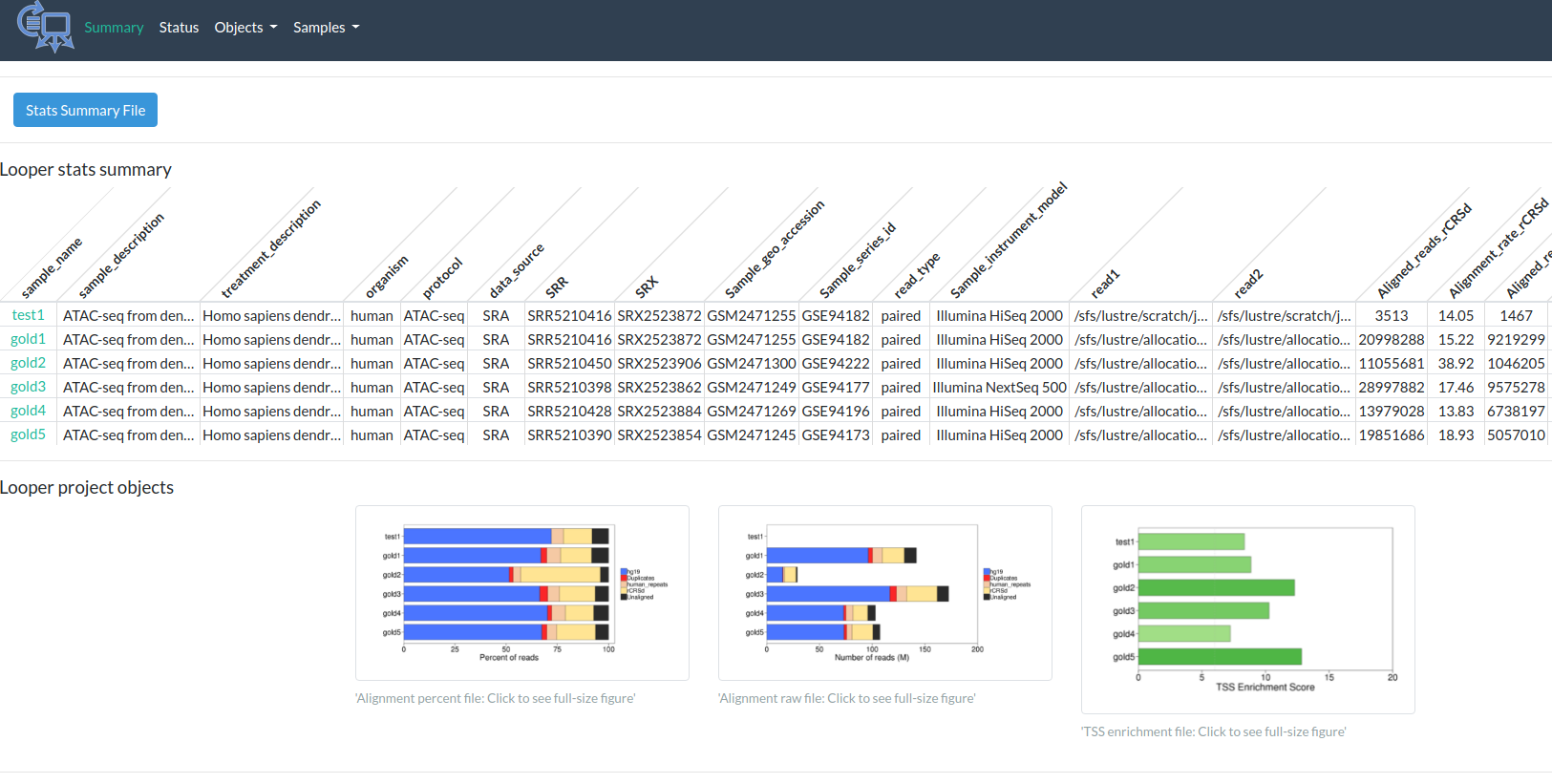
http://pepatac.databio.org/en/latest/files/examples/gold/gold_summary.html
PEPATAC in practice
Conclusion
Everyone else
Eat chicken nuggets!

Eat chicken nuggets!
Thank You
Sheffield lab
John Lawson
Vince Reuter
Jason Smith
Jianglin Feng
Michal Stolarczyk
Aaron Gu
Christoph Bock
Andre Rendeiro
Johanna Klughammer
Howard Chang
Ryan Corces
Yuning Wei
Jin Xu
Funding:






Parallelism Philosophy

by process


by sample


by dependence


Very easy
Easy
Hard




