Epigenome tools: ATAC-seq and Bisulfite-seq
Nathan Sheffield, PhD

What is epigenomics?
Epigenomics is the study of the physical modifications, associations and conformations of genomic DNA sequences (Schwartzman and Tanay 2015)
Epigenomics is the study of the chemical modification and physical conformation of cellular DNA and bound proteins
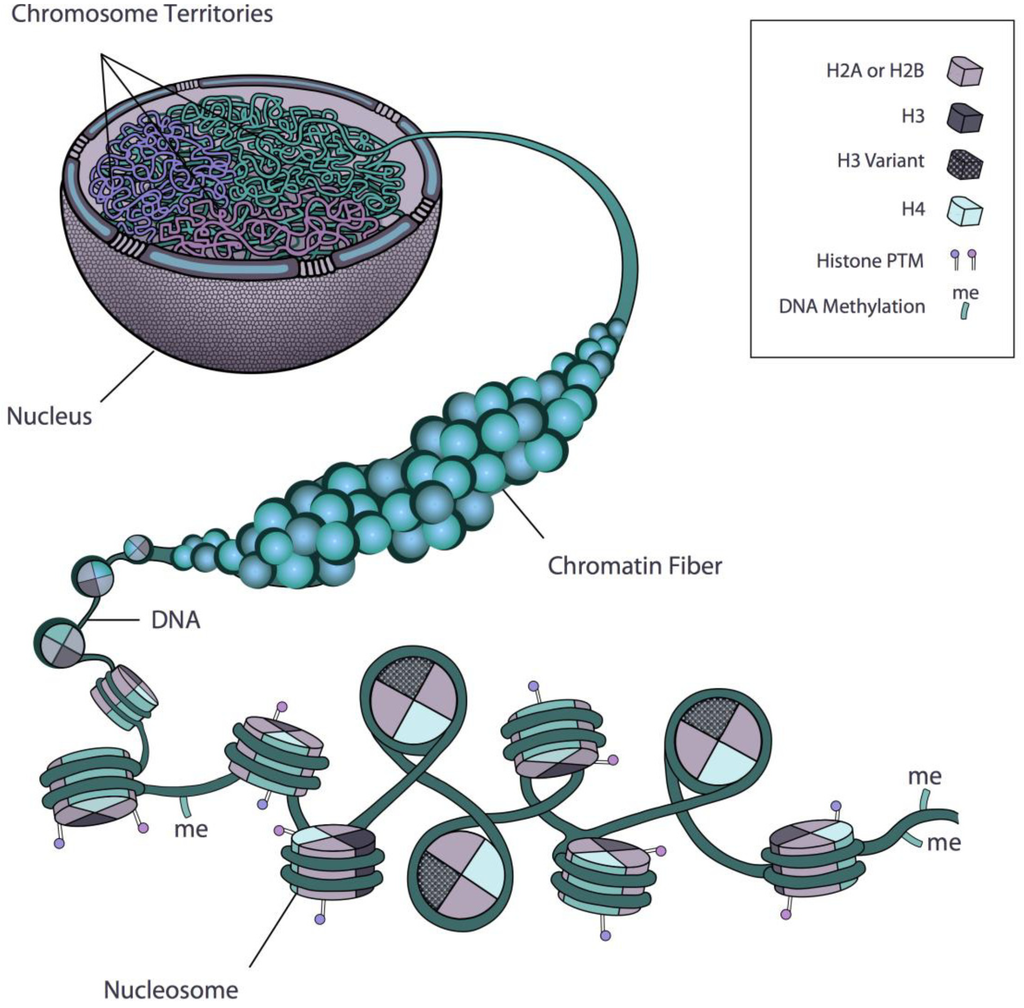
Rosa and Shaw 2013
The epigenome

If we can measure how DNA is packaged,
we can understand what a cell is doing
Epigenomics
the study of the chemical modification and physical conformation of cellular DNA and bound proteinsEpigenetics
???What is epigenetics?
the causal study of embryological development (Waddington 1957, The strategy of the genes)
The study of mitotically and/or meiotically heritable changes in gene function that cannot be explained by changes in DNA sequence
(Riggs et al. 1996)
(Riggs et al. 1996)
a change in the state of expression of a gene that does not involve a mutation, but that is nevertheless inherited in the absence of the signal (or event) that initiated the change. (Ptashne and Gant 2002)
What is epigenetics?
the structural adaptation of chromosomal regions so as to register, signal or perpetuate altered activity states. (Bird 2007)
Epigenetics refers to changes in gene regulation brought about through modifications to the DNA's packaging proteins or the DNA molecules themselves without changing the underlying sequence.
(Lord and Cruchaga 2014, Nature Neuroscience)
(Lord and Cruchaga 2014, Nature Neuroscience)
the study of the mechanisms that allow cells to translate the nearly constant genome content of a multicellular organism into multiple functional and stable cellular conditions (Schwartzman and Tanay 2015)
What is epigenetics?
The word literally means "on top of genetics," and it's the study of how individual genes can be activated or deactivated by life experiences. (The Week, 2013)

Epigenomics
the study of the chemical modification and physical conformation of cellular DNA and bound proteinsEpigenetics
???What does the genome encode?
Chromatin accessibility
ATAC-seqDNA methylation
Bisulfite-seqWhat is regulatory DNA?

Regulatory DNA is a decision-maker
Challenges to studying regulatory DNA
- Variation: age, cell-type, environment, disease
- Amount: 1-2% protein coding vs 8-20%? regulatory
- Target: what gene does it affect?
- Function: is it a promoter, silencer, insulator, enhancer?
- Rigidity: genetic code vs TF motifs
Genetic code
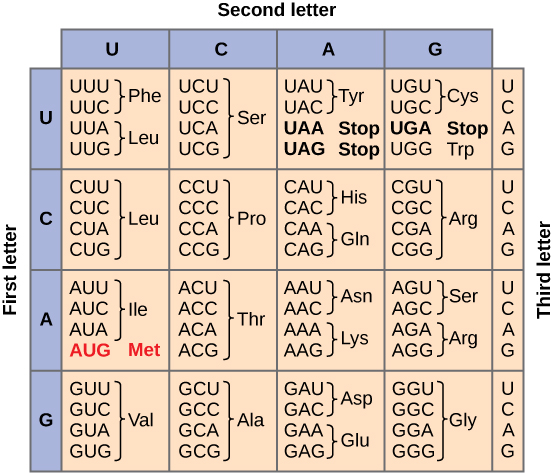

Transcription factor motif

We can computationally identify genes and even predict function. Regulatory DNA is more difficult.

Chromatin accessibility
Chromatin accessibility is the degree to which nuclear macromolecules are able to physically contact chromatinized DNA...
[It] is determined by the occupancy and topological organization of nucleosomes as well as other chromatin-binding factors that occlude access to DNA.
Klemm et al. 2019
How can we identify regulatory DNA?
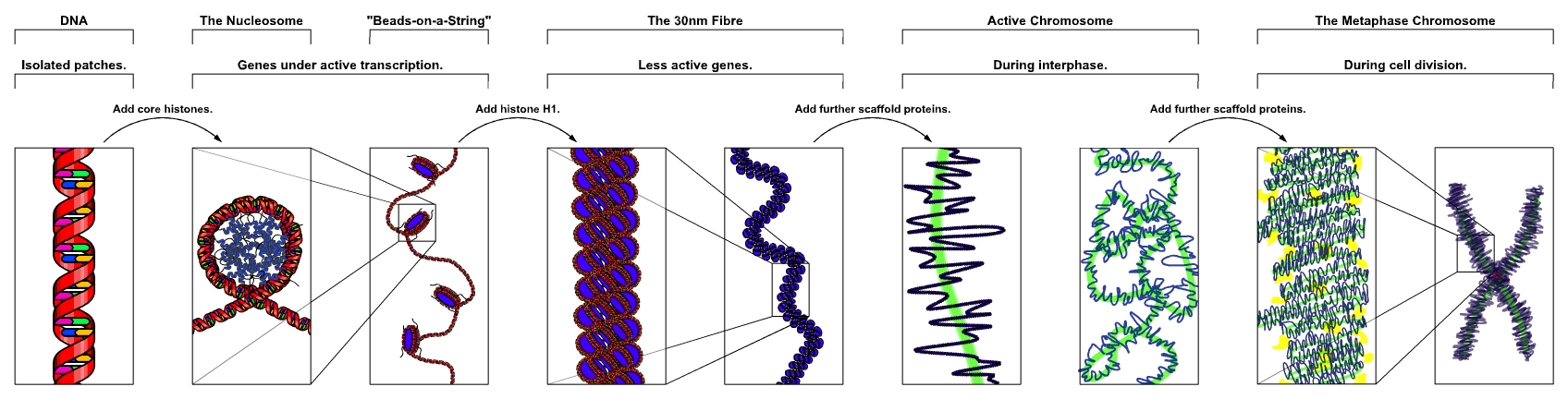

https://en.wikipedia.org/wiki/Chromatin
How can we identify regulatory DNA?
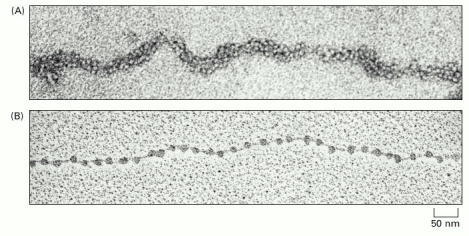
Alberts 2002
How can we identify regulatory DNA?
- ChIP: Chromatin immunoprecipitation
- DNase: classic 'gold standard' to identify open chromatin
- ATAC: Assay for transposase-accessible chromatin
- FAIRE: Formaldehyde-assisted isolation of regulatory elements
ChIP-seq
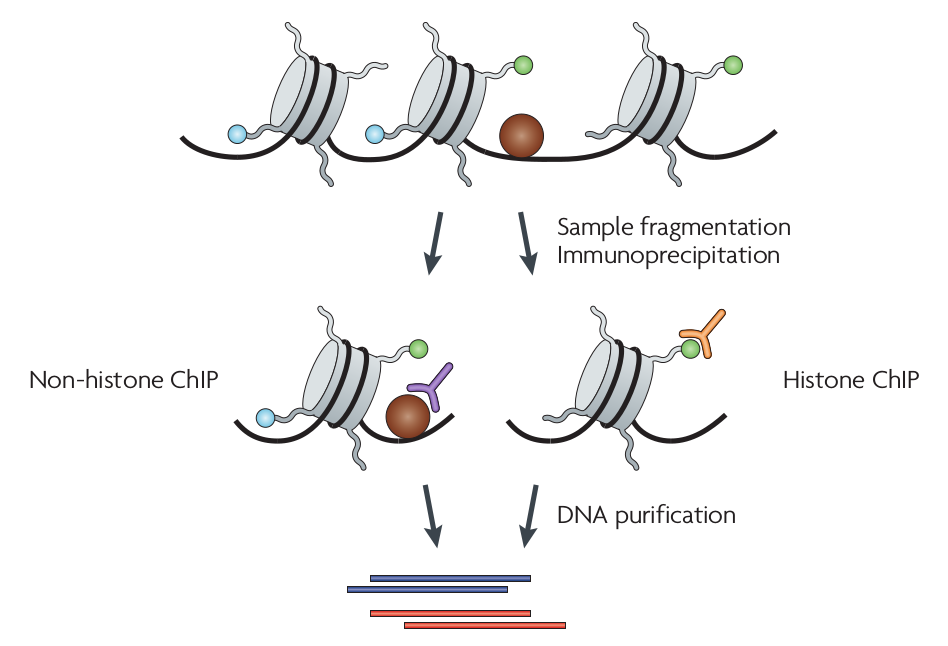
DNase-seq: Biology

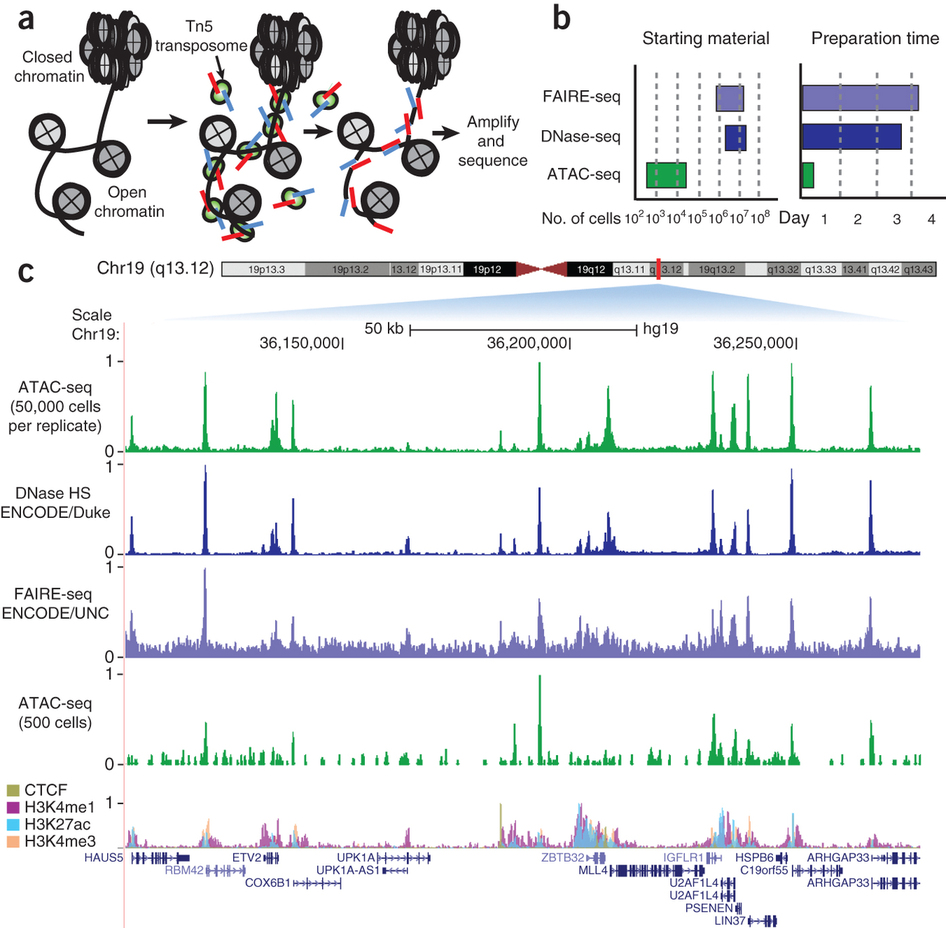
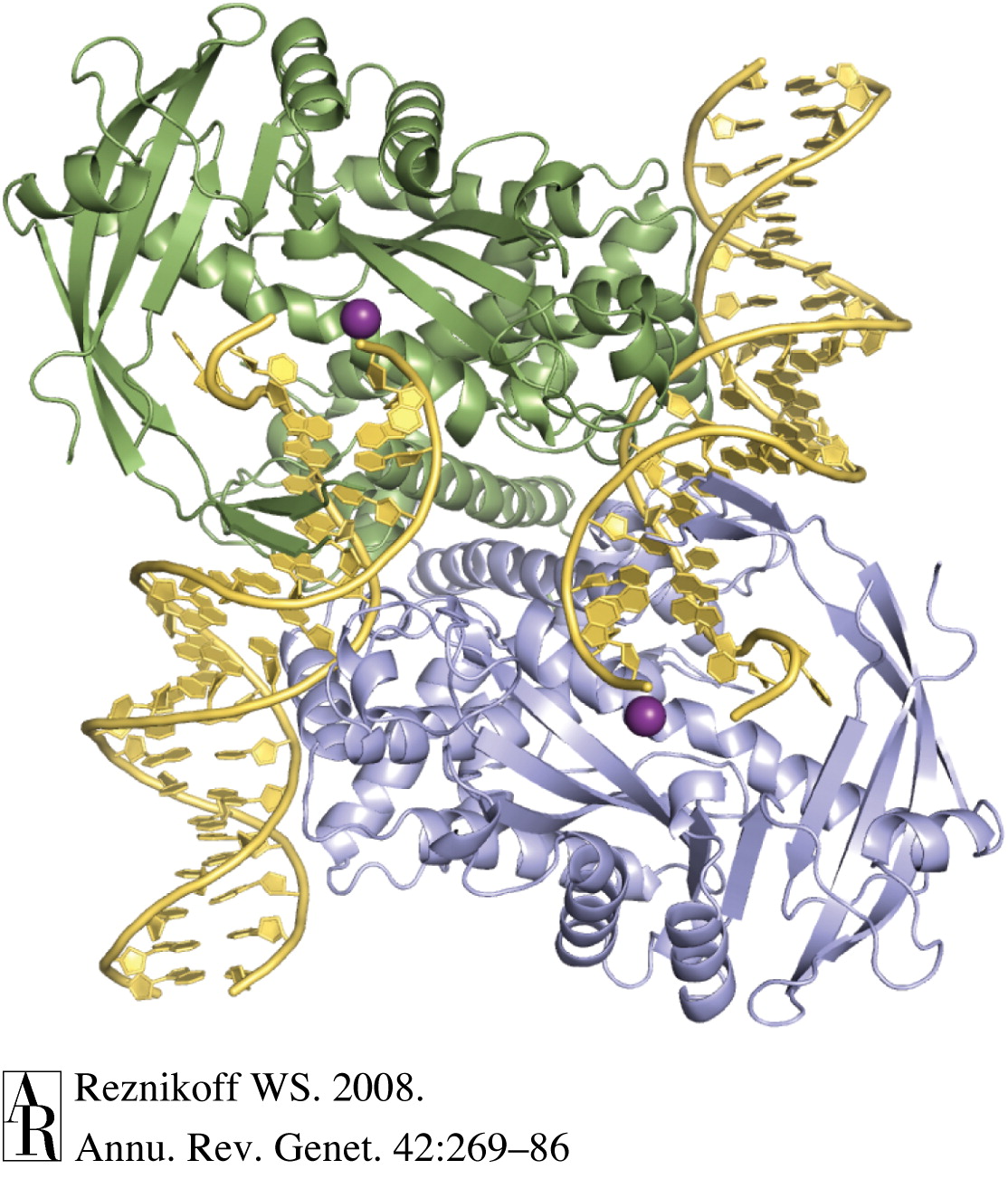
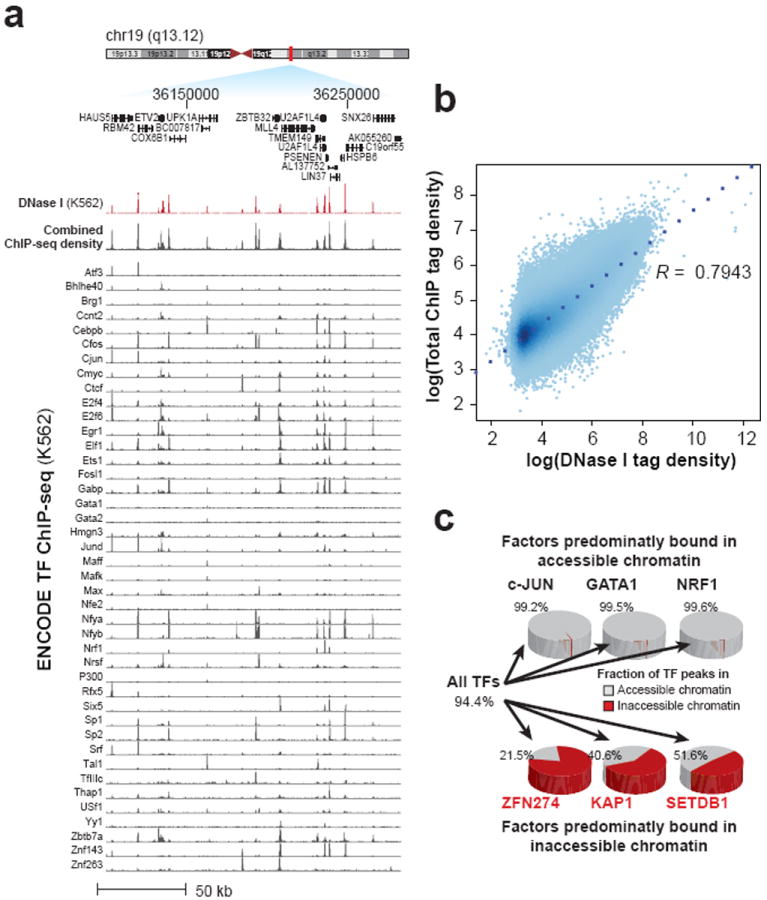
Chromatin accessibility biology summary
- Open chromatin usually coincides with active regulatory DNA
- ... but exact annotation or binding is not provided
- The advantage and disadvantage of ChIP seq is in its target. It also requires antibodies and provides more diffuse signal
- ATAC is pronounced 'attack'
Basic data analysis steps
- 1. Trim adapters (fastq -> fastq)
- 2. Align reads (fastq -> bam)
- 3. Shift reads (bam -> bam)
- 4. Call peaks (bam -> bed)
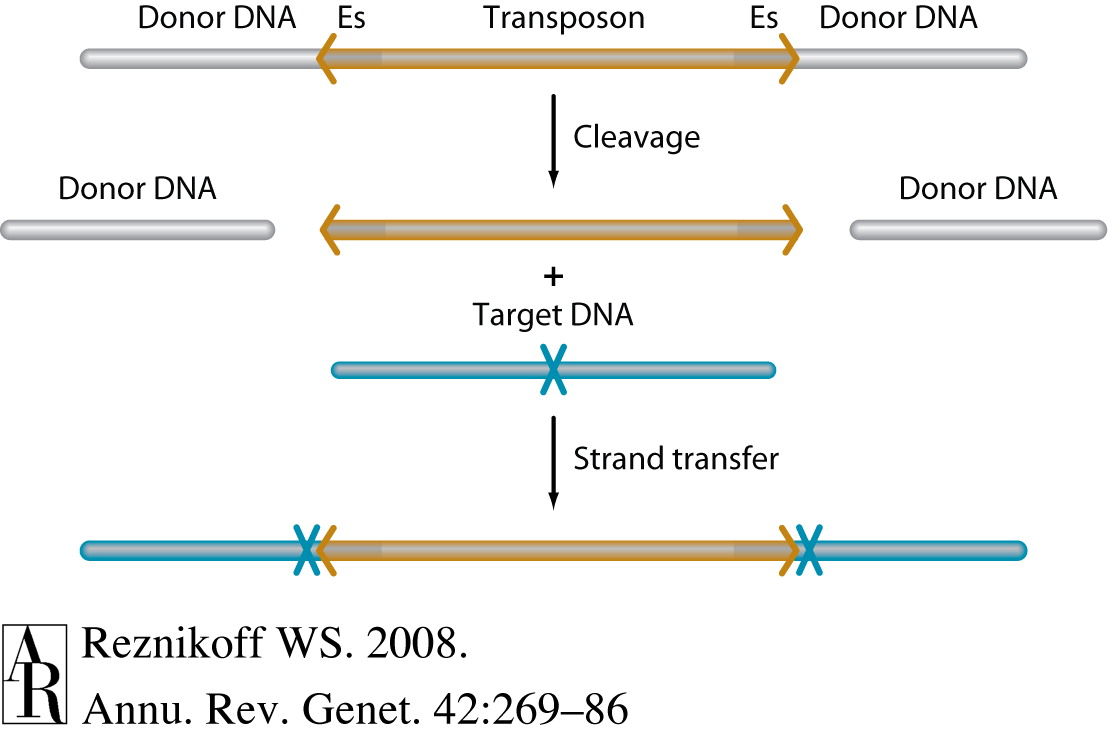
Tn5 molecular biology

What is DNA methylation?

DNA methylation: distribution
- Cytidine and Adenine can be methylated
- In mammals, DNA methylation happens at CpG dinucleotides
- In mammals, DNA methylation rarely happens at CHH dinucleotides (H= A/G/T)
DNA methylation: evolution and development
- Plants have cytidine and adenine methylation
- Yeast is typically unmethylated (but some species have some)
- In mice, DNMT knockouts are embryonic lethal
DNA methylation: heritability
- The classic "epigenetic" mark
- Heritability: DNA Methyltransferases (DNMTs) copy from parent to child strand
- Maintenance methyltransferases: DNMT1
- De novo methyltransferase: DNMT3(a/b)
DNA methylation: hemimethylation

DNA methylation: imprinting
- "Imprinted" regions are parent-specific, binary allele-specific methylation
- Classic example: X-inactivation. DNA methylation is critical for maintenance but not establishment of the silent X.
DNA methylation: signal levels
- An individual cytosine is either methylated or not methylated
- Continuous measures arise via averaging at multiple resolutions: strands, alleles, cells, cell-types and genomic regions
DNA methylation: gene regulation
- Many (not all) transcription factors are methylation-sensitive

DNA methylation in cancer
- Most cancers have globally decreased methylation with punctate increased methylation
- azacytidine is a cytidine analog used in cancer treatment.

DNA demethylation
In the active case, appears to happen via hydroxymethylation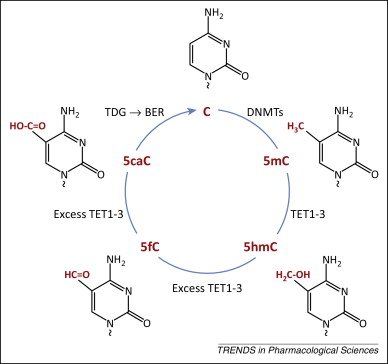
Ivanov et al. 2014
DNA methylation: measuring
- MeDIP assays rely on methylation-sensitive antibodies
- bisulfite microarrays use base-pair hybridization
- bisulfite sequencing is the de facto standard
 Bisulfite-seq
Bisulfite-seq
 Bisulfite-seq
Bisulfite-seq

CTGACTGTCGATCGATCGGATCATAGTCAGCTAGCATTTTGGGACCGCG - Reference genome | | | | TTGATTGTTGATTGATTGGATTATAGTTAGTTAGTATTTTGGGATCGCG - Bisulfite-converted
How can you align this sequence? Convert the reference!
TTGATTGTTGATTGATTGGATTATAGTTAGTTAGTATTTTGGGATTGTG - Converted reference | | | | TTGATTGTTGATTGATTGGATTATAGTTAGTTAGTATTTTGGGATTGTG - Fully converted
RRBS: Reduced Representation Bisulfite Sequencing

(Baheti et al. 2016)


Advanced analysis concepts



Locus Overlap Analysis












Slides at http://databio.org/slides/epigenome_tools.html